Brain Imaging Generation with Latent Diffusion Models
Making Brains Out of Thin Air (And lots of MRI scans) - as presented in the seminar offered by DKFZ Heidelberg WS24/25. Full presentation link available on my website.
Imagine if we could generate high-resolution MRI scans of the human brain—synthetic images that look just like real MRIs but without the hassle of patient data privacy issues or limited sample sizes. That’s exactly what Latent Diffusion Models (LDMs) are bringing to the table. Let’s break it down!
This work is based on the research by Pinaya et al. (2022) "Brain Imaging Generation with Latent Diffusion Models".
This presentation explores the generation of high-resolution MRI brain scans using Latent Diffusion Models (LDMs). The study addresses challenges in medical imaging, where data scarcity and privacy concerns limit access to diverse datasets. By leveraging generative networks (Latent Diffusion Models), the study synthesizes realistic MRI scans conditioned on age, sex, and brain structure volumes.
Key Takeaways
- LDMs offer a novel approach to generating synthetic MRI scans
- DDIM significantly improves training efficiency (25x faster)
- The model achieves state-of-the-art results while maintaining diversity
- Generated images show promising potential for medical research and training
- Generated dataset (100 000 samples) publicly released
Motivation
Medical imaging is a goldmine for AI, but there’s one major issue: data is scarce. This is due to three key factors:
- Privacy Concerns: GDPR and other regulations make sharing real patient data particularly challenging
- Rare Pathologies: Some diseases appear too infrequently in datasets to build robust models
- Data Complexity: MRI scans are 3D and high-resolution, making them significantly more complex than standard 2D images
Additionally, Deep Neural Networks (DNNs) are data-hungry—they require vast amounts of data to perform well.

To deal with this, researchers turned to generative models—AI models that can synthesize new data.
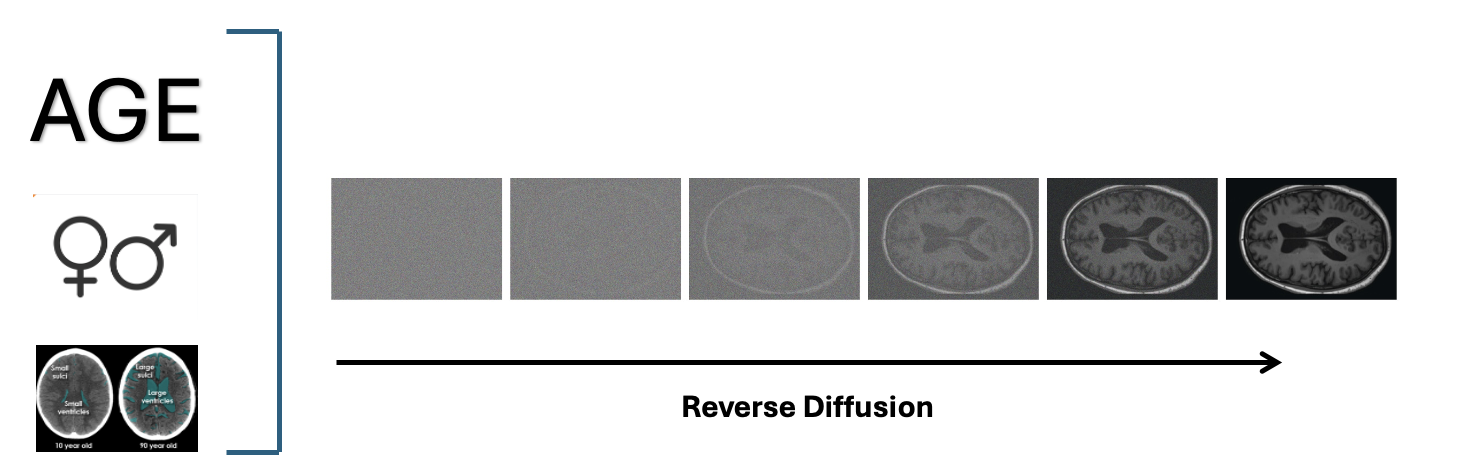
Challenges
Reading MRIs is complex as it is, but there are other factors which impact the specialist’s ability to objectively assess MRI data. The human visual apparatus is the result of millions of years of evolution and can do some incredible things, but noticing slight changes in curvature, color luminance or accurately estimating area was never one of them. Turning to AI could perhaps one day overcome these inherent limitations all humans possess by virtue of being simply that, humans.
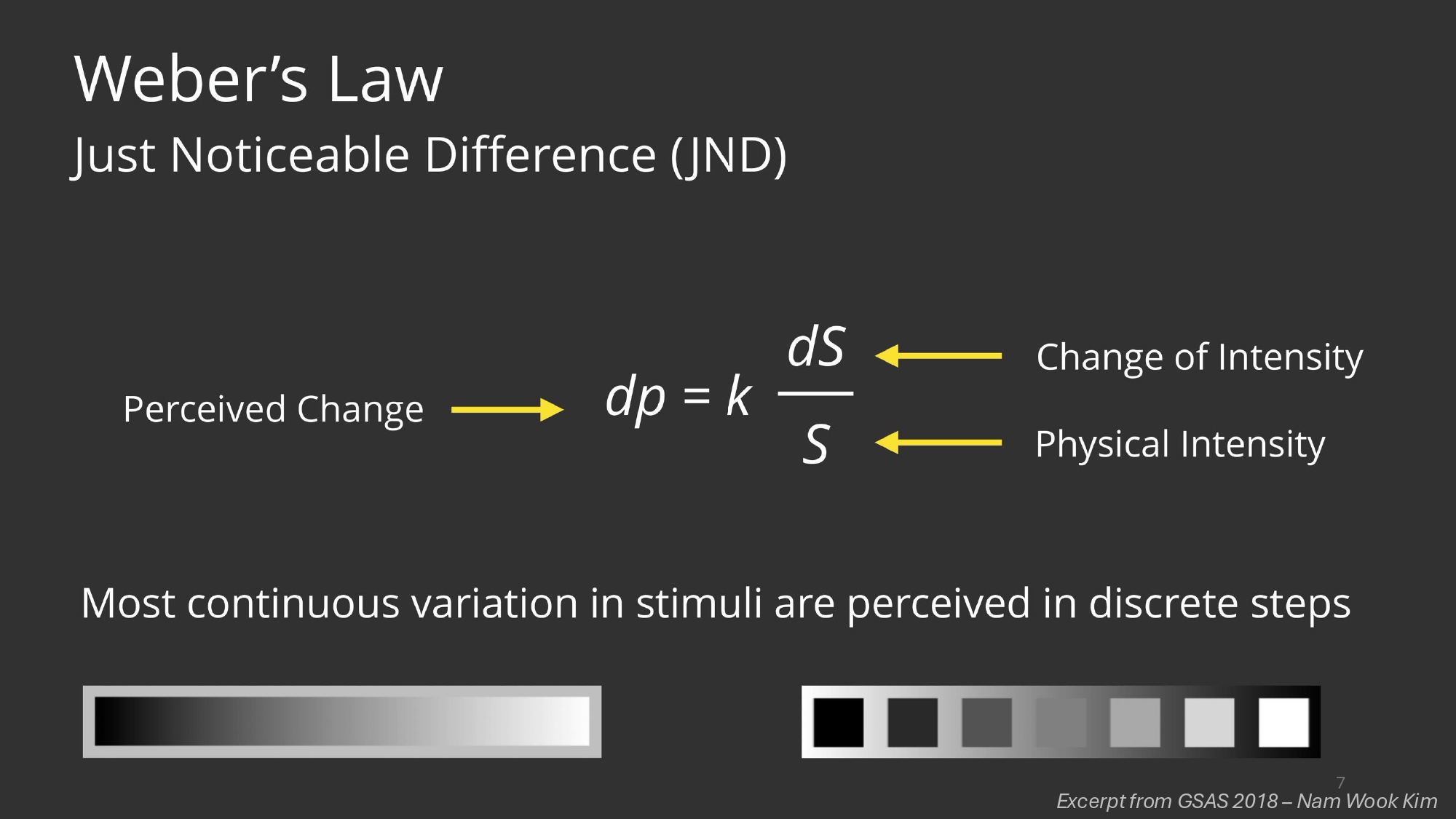

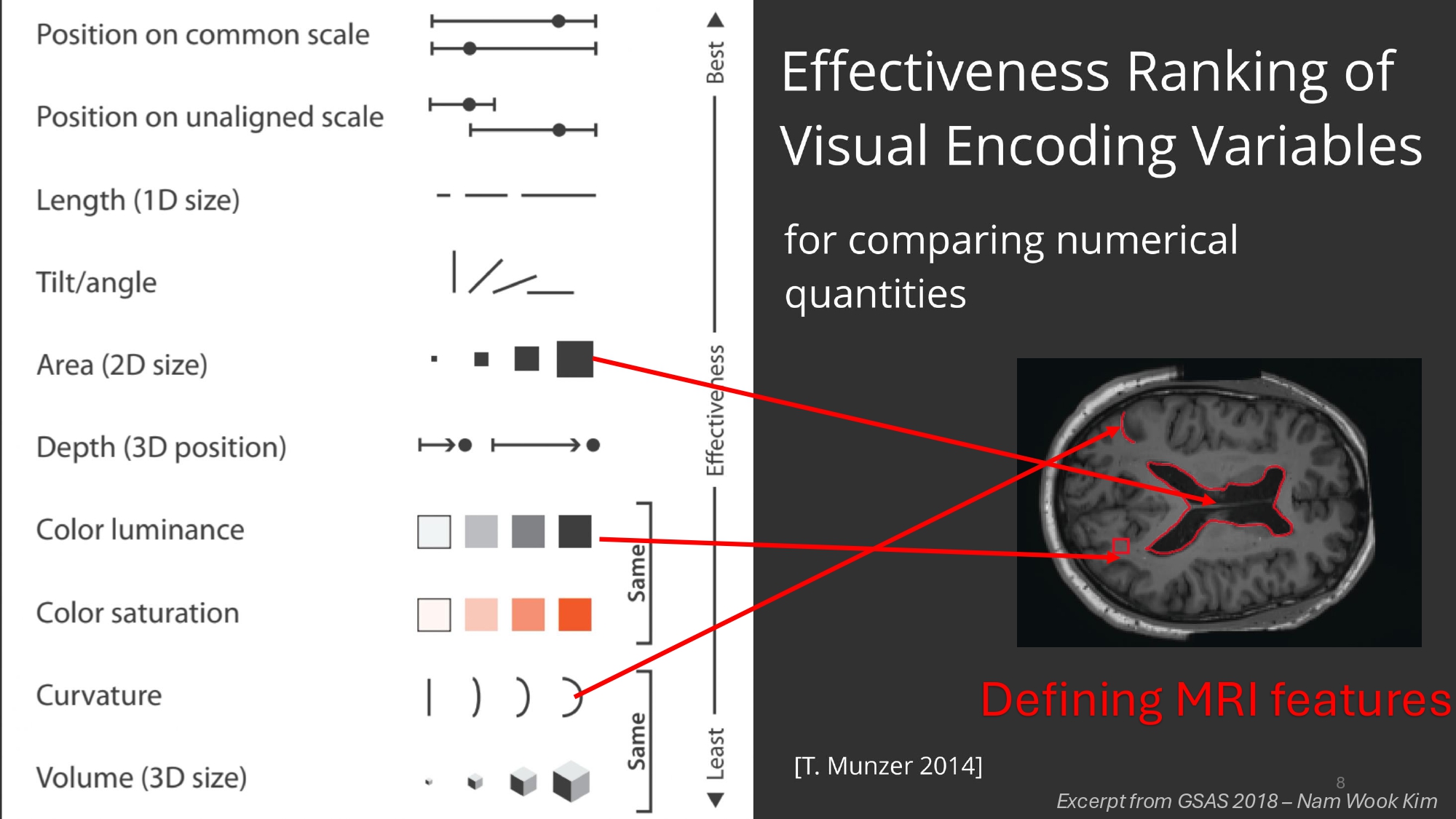
How complex is actually MRI data? To make things even more complex, whereas in other downstream computer vision tasks individual pixels alone don’t count for much, in this case a few voxels make the difference between a deadly pathology and a healthy individual (this is the brain we’re talking about after all) so high resolution is really important.

Key Takeaways
- Perceptual Limitations: Humans struggle with:
- Noticing slight changes in curvature
- Accurately estimating areas
- Detecting subtle intensity variations in grayscale images
- Data Complexity: MRI data is particularly challenging because:
- Individual voxels can be crucial for diagnosis
- High resolution is essential for accurate assessment
- 3D structure requires complex analysis
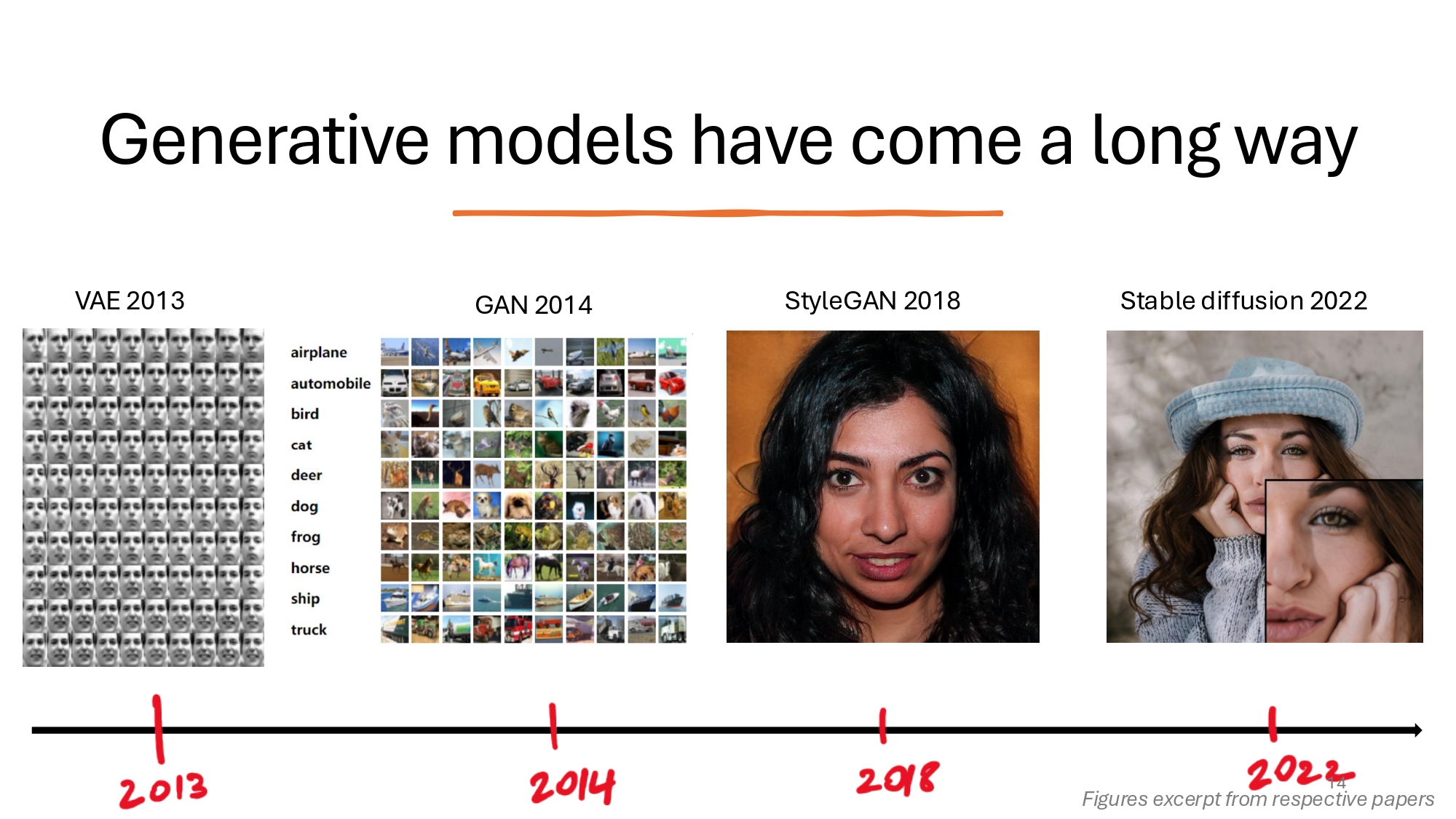
The Evolution of Generative Models
The field of generative AI has evolved dramatically since its early days. Starting with Variational Autoencoders (VAEs) in 2013, which excelled at learning latent representations but produced notoriously blurry outputs, the field soon saw the emergence of Generative Adversarial Networks (GANs) in 2014. GANs represented a major breakthrough in AI-generated images, though they frequently encountered issues with mode collapse. The real revolution came in 2022 with Stable Diffusion, which leveraged diffusion-based approaches to take image generation capabilities to unprecedented heights. This evolution set the stage for Latent Diffusion Models (LDMs), which would go on to fundamentally transform the landscape of generative AI.
Why Latent Diffusion?
Traditional Diffusion Models add noise to an image and then train a network to recover it. But this process in pixel space is inefficient. How inefficient exactly?
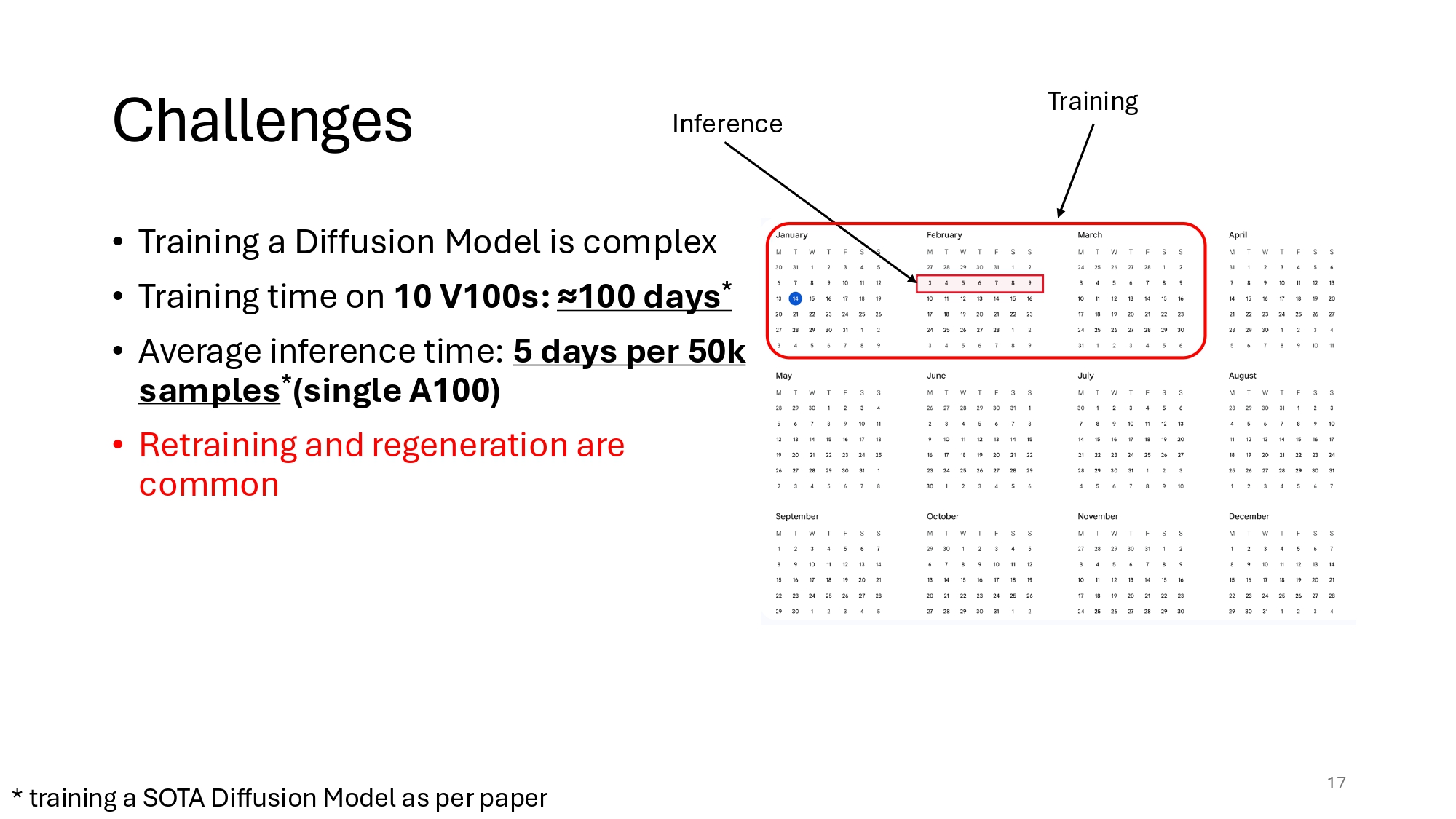
LDMs make use of a simple observation, that is not even new: the pixel space is overly redundant and adds unnecessary dimensionality. We know from early Signal Processing that the high frequency of images (if we decompose into a Fourier or Cosinus transform) corresponds to imperceptible details, because of the inherent spatial biases of the pixel space:
-
Nearby pixels in an image are often correlated, as they belong to the same object or surface. This spatial coherence means that patterns like edges or textres are typically local phenomena.
-
Objects in images retain their identity and characteristics regardless of their position. A cat remains a cat whether it’s in the top-left or bottom-right of the image.
-
High-level patterns (e.g., objects) in images are composed of lower-level features (e.g., edges, corners, textures).
-
Objects in natural images tend to have well-defined boundaries and shapes, often forming closed contours.
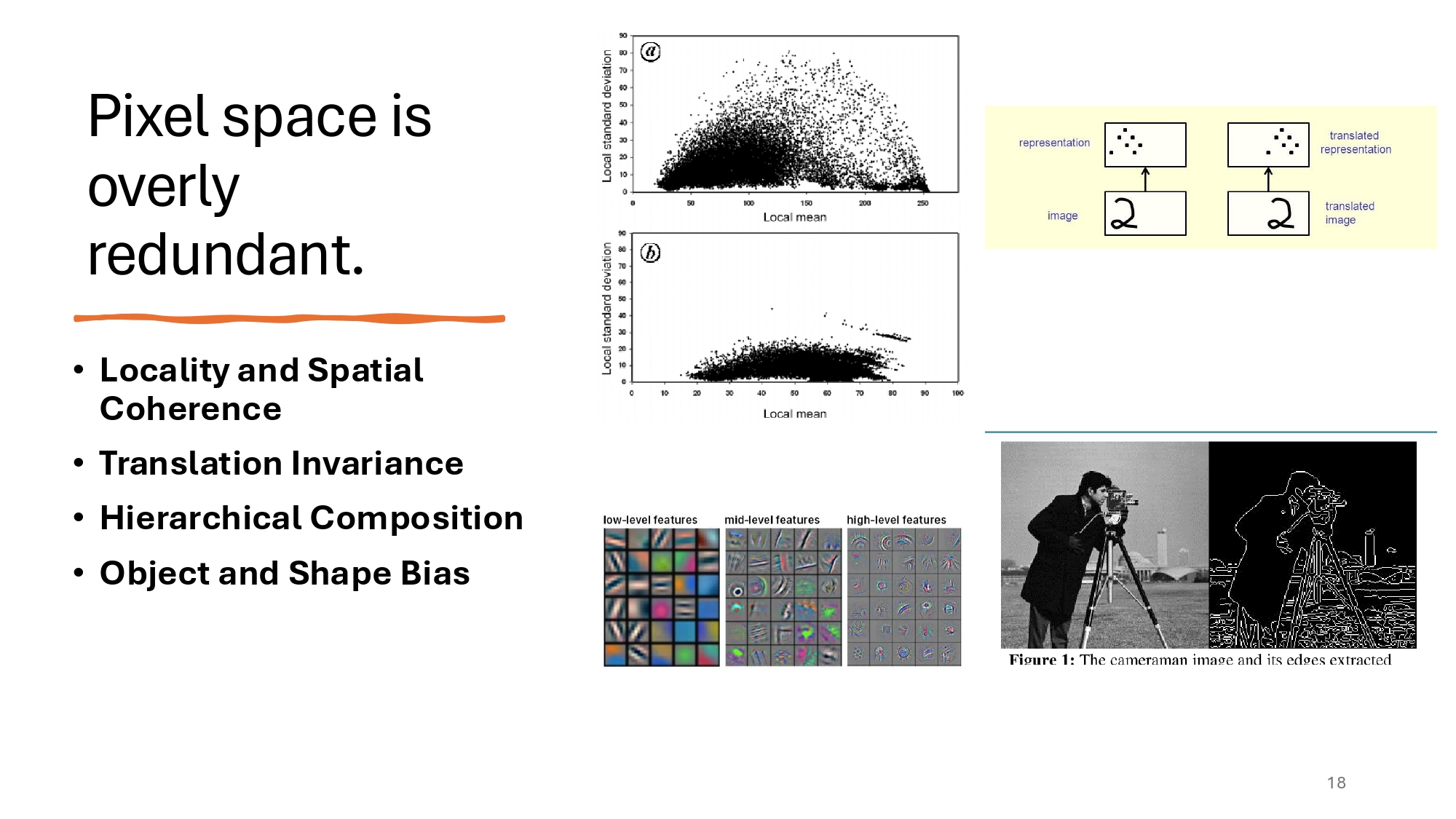
LDMs solve this by operating in latent space instead. This approach reduces computational redundancy since pixel space contains too much detail, encodes images more efficiently by keeping only the most important features, and allows conditioning on variables like age, sex, and brain structure volumes. By working in latent space rather than pixel space, LDMs achieve better results with less computational overhead.
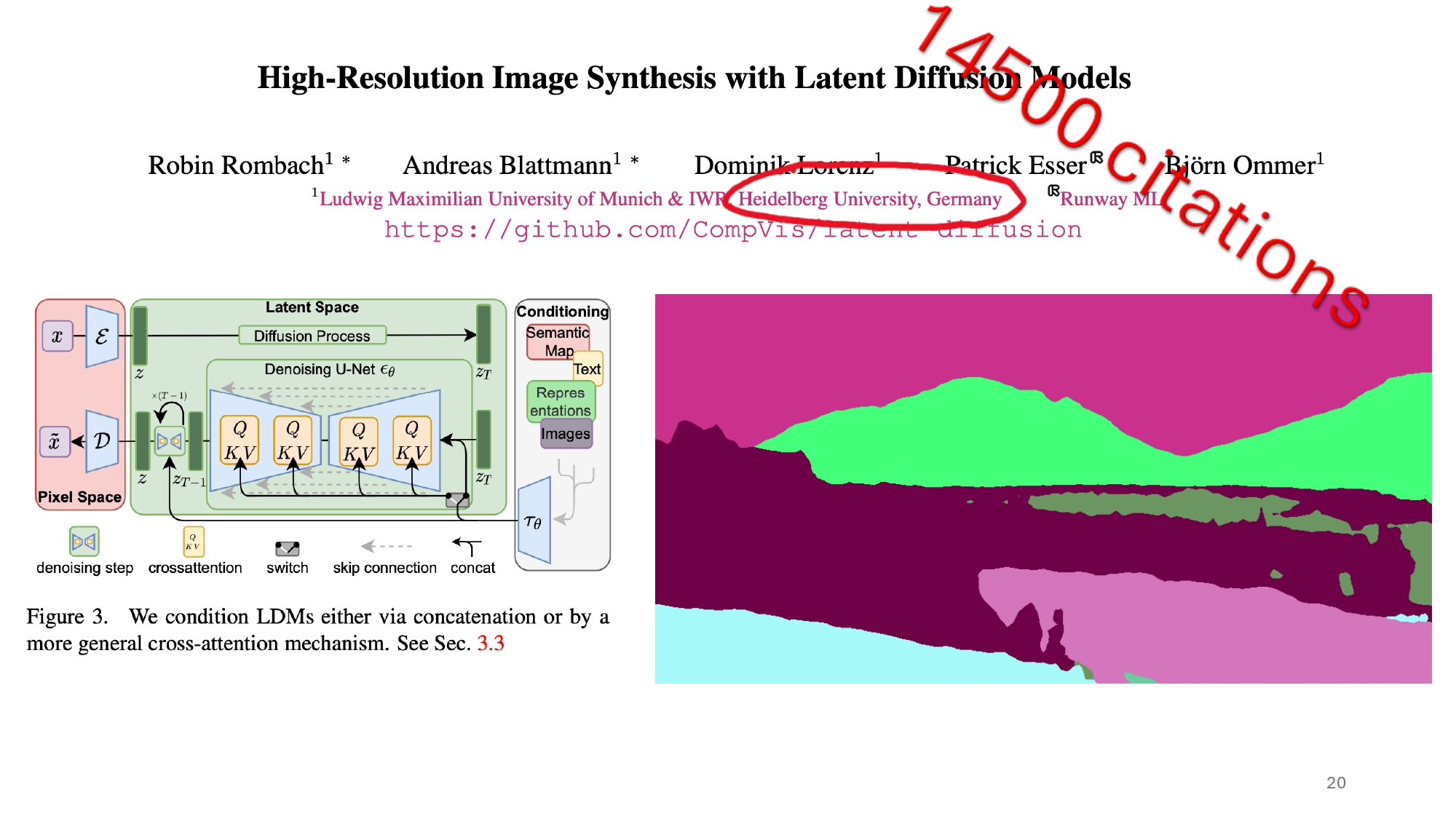
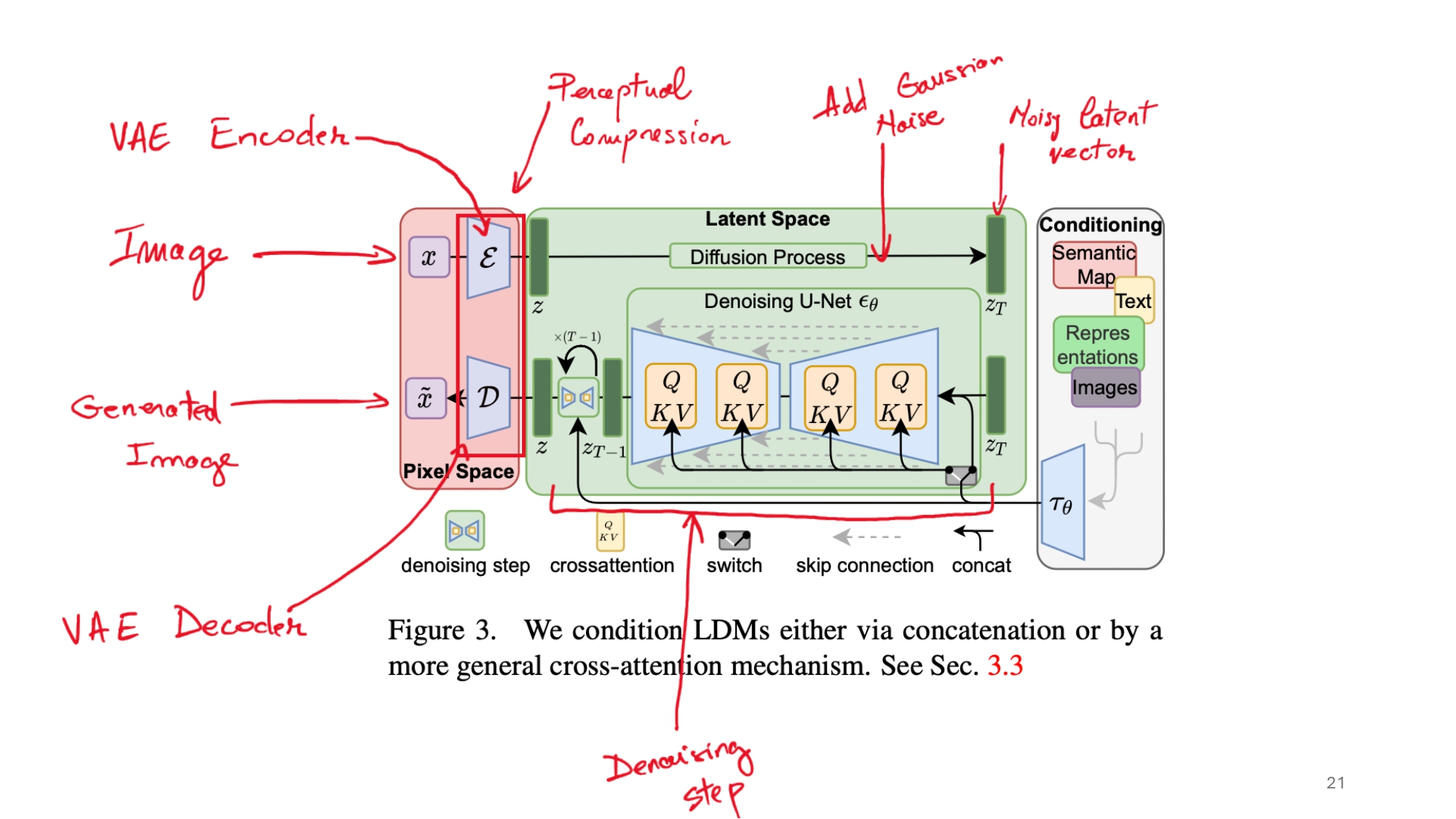
LDM Architecture Deep Dive
The architecture of Latent Diffusion Models is quite sophisticated, combining several key components:
-
The Autoencoder: First, a VAE compresses the high-dimensional brain MRI images into a lower-dimensional latent space. This compression significantly reduces the computational overhead of the diffusion process.
- The U-Net Backbone: At the heart of the LDM is a U-Net architecture that learns to denoise images. The U-Net consists of:
- An encoder path that downsamples features
- A decoder path that upsamples back to the original resolution
- Skip connections that preserve fine details
- Residual blocks for better gradient flow
- Self-attention layers for capturing long-range dependencies
- Conditioning Mechanisms: The model incorporates multiple conditioning signals:
- Cross-attention layers inject demographic information (age, sex)
- AdaIN (Adaptive Instance Normalization) layers incorporate brain structure volumes
- These conditions help guide the generation process towards specific desired attributes
- The Diffusion Process: In the latent space, the model:
- Gradually adds noise to latent representations
- Learns to reverse this process step by step
- Uses timestep embeddings to track the denoising progress
So LDMs should make everything faster, right? Well, not exactly:
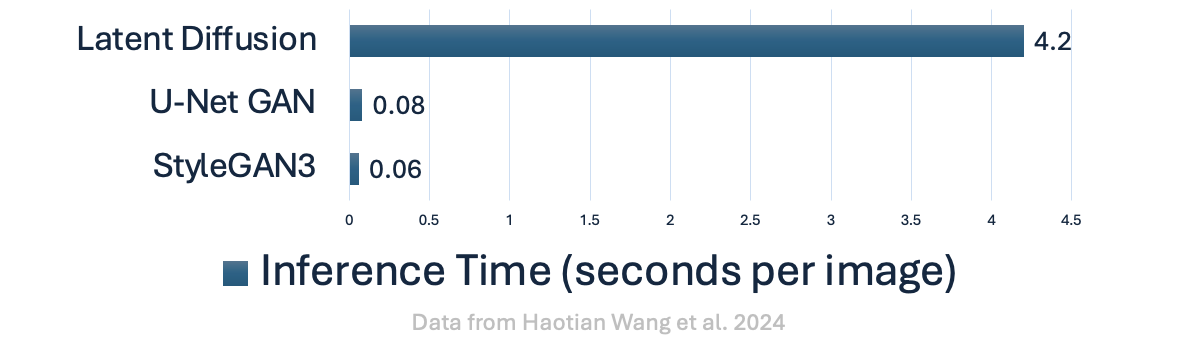
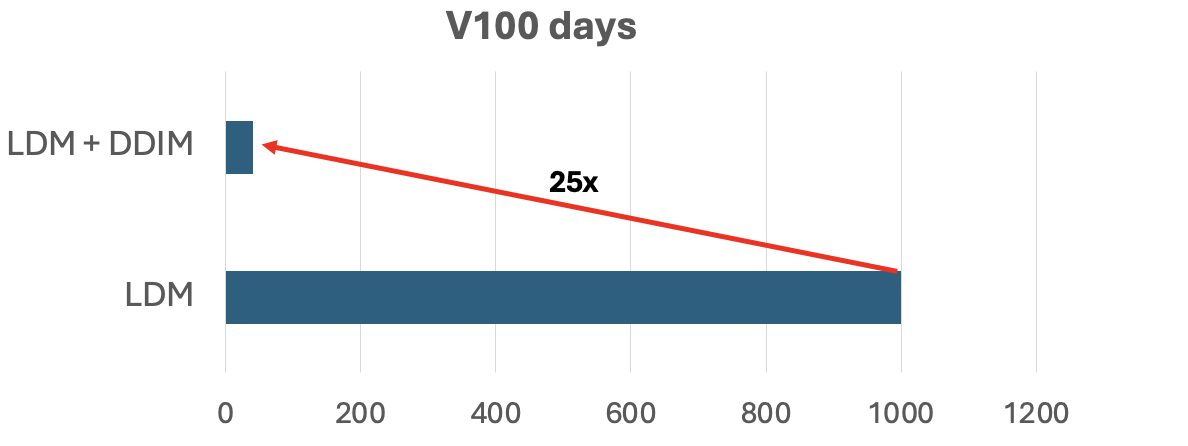
Denoising Diffusion Implicit Models (DDIM)
One major issue with Diffusion Models (and LDMs for that matter) is their slow sampling speed. Enter DDIM, which replaces the traditional stochastic sampling with a non-Markovian deterministic update. This speeds up training 25x, cutting down training time from 1000 GPU days to just 40! The key idea is to replace the stochastic forward process with a deterministic one. Given a clean image \(x_0\), the forward process in DDIM is defined as:
\[x_t = \sqrt{\bar{\alpha}_t}x_0 + \sqrt{1-\bar{\alpha}_t}\epsilon\]where:
- \(x_t\) is the noisy image at timestep t
- \(\bar{\alpha}_t\) represents the cumulative product of noise scheduling
- \(\epsilon\) is random Gaussian noise
The reverse process (denoising) then becomes:
\[x_{t-1} = \sqrt{\bar{\alpha}_{t-1}}(\frac{x_t - \sqrt{1-\bar{\alpha}_t}\epsilon_\theta(x_t,t)}{\sqrt{\bar{\alpha}_t}}) + \sqrt{1-\bar{\alpha}_{t-1}}\epsilon_\theta(x_t,t)\]This deterministic formulation allows for:
- Fewer sampling steps (50 vs 1000)
- Consistent high-quality outputs
- Significantly faster training and inference
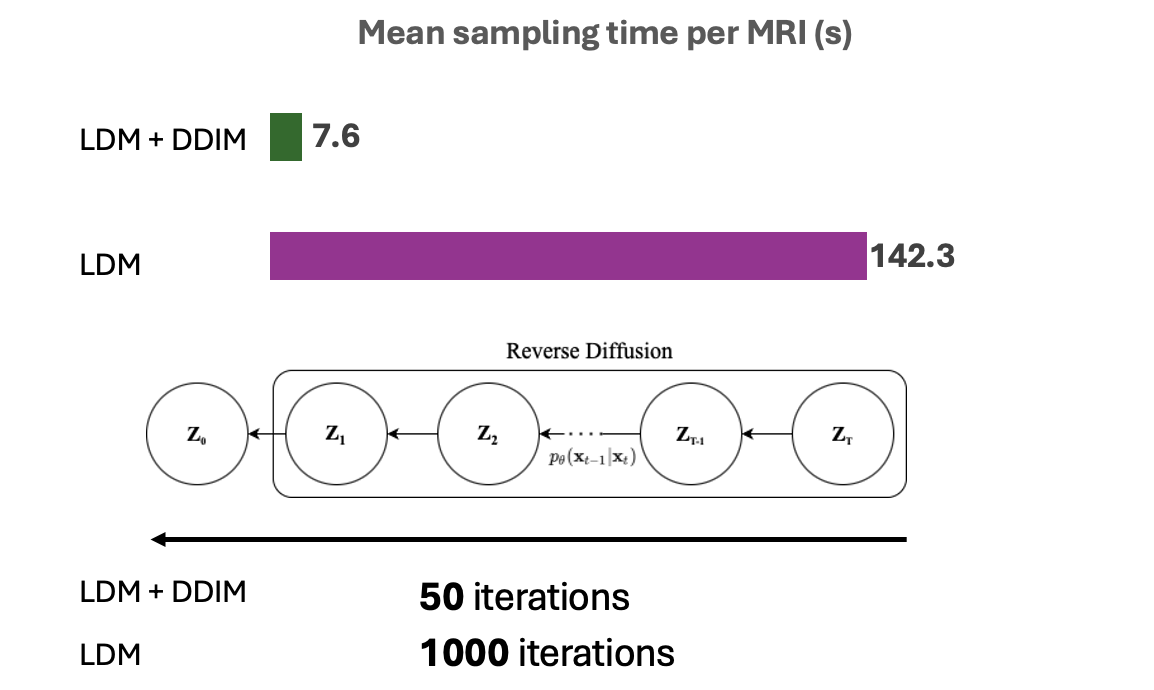
Results
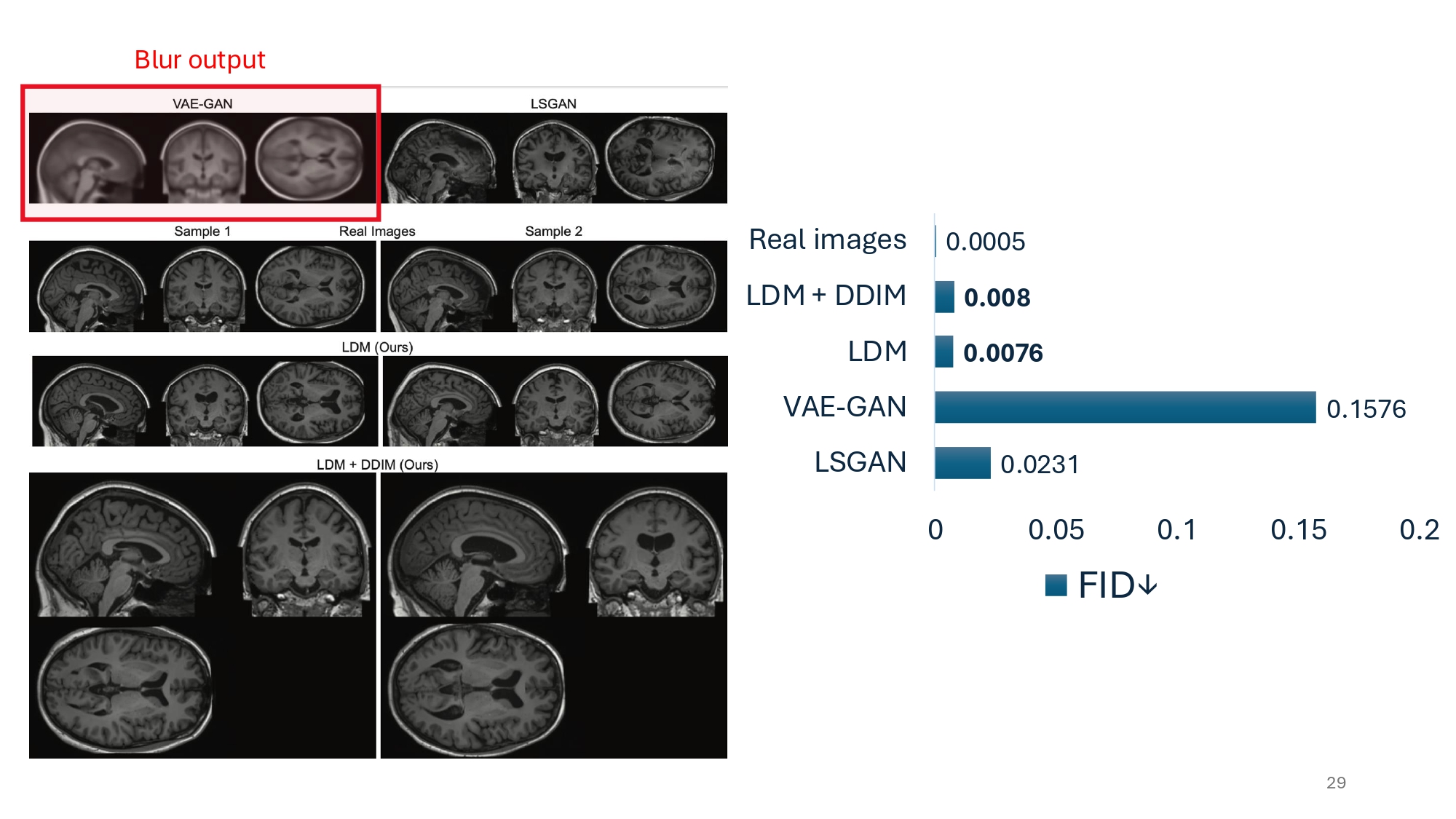
Their results are really impressive. The generated image quality, albeit being 15 times worse than the real images, it is still a staggering 21 times better than the former state-of-the-art (using VAE-GAN). It is important to note how the authors use their metrics.
The Fréchet Inception Distance (FID) measures how similar generated images are to real ones. It works by comparing feature distributions extracted from a pre-trained neural network (Inception v3). Generated images are considered high quality when they have a low FID score, meaning their features closely match those of real images. Instead of an Inception v3, which would not make much sense here, the authors use a Med3D network to extract informative features.
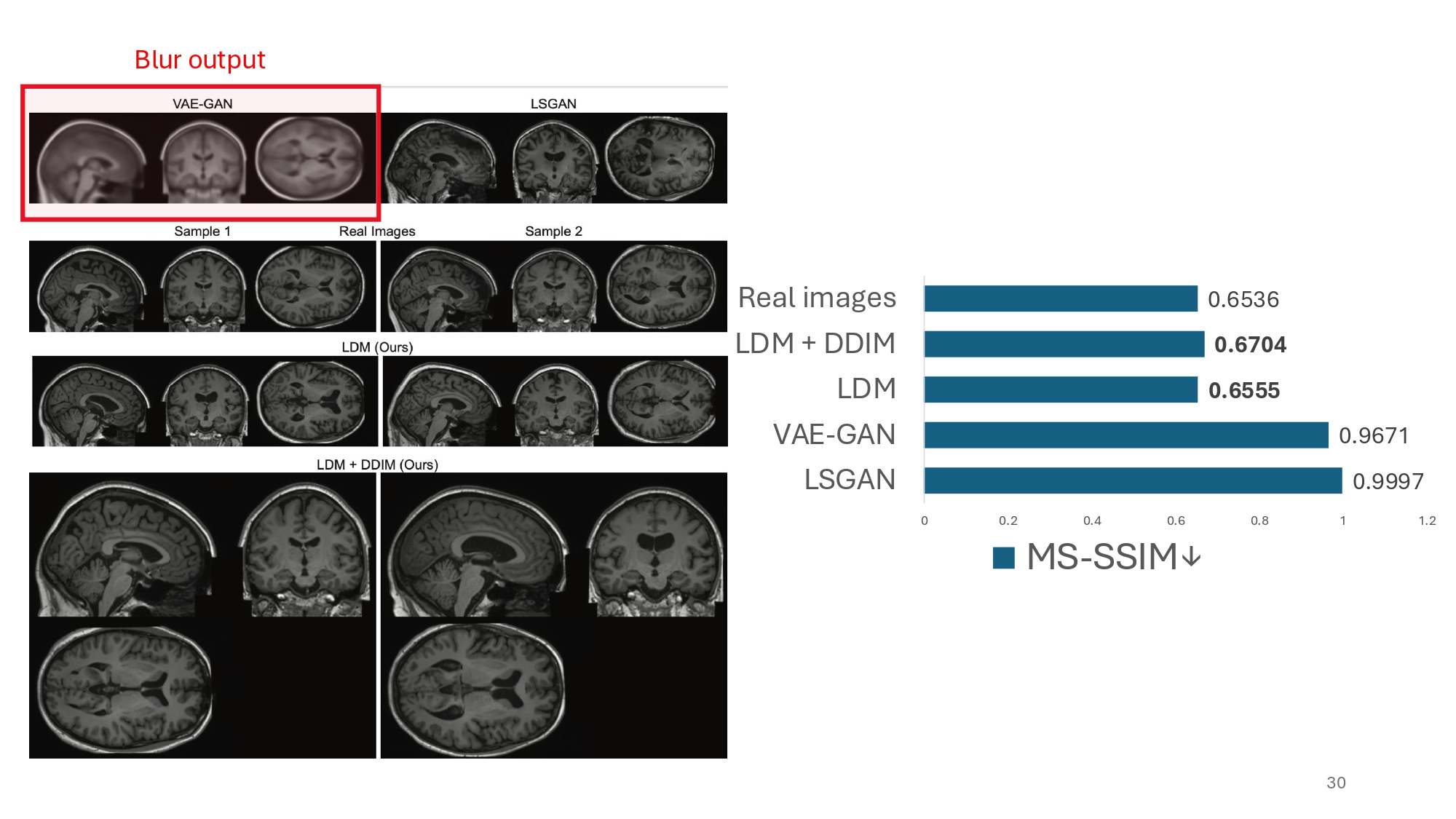
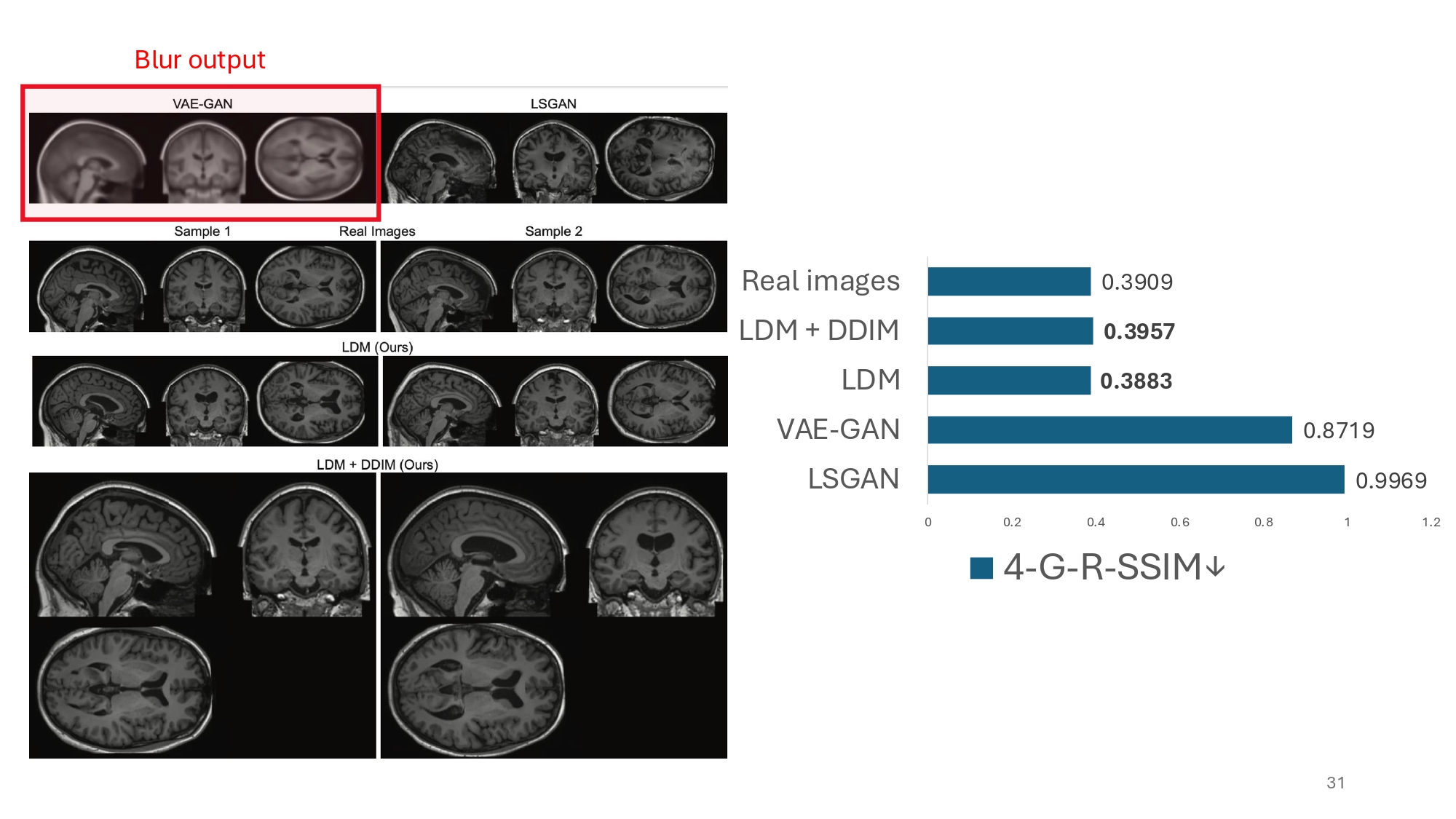
What is also interesting to observe is that authors use two variants of SSIM, a metric traditionally designed for measuring similarity between image pairs, to assess diversity over the generated images. In terms of diversity the authors were able to obtain on par results with the real world images. That is quite an achievement.
Dataset
Here is a quick description of the dataset they have used.
| Dataset Property | Description |
|---|---|
| Image Type | T1-weighted MRI images sourced from the UK Biobank (UKB) dataset |
| Population | Healthy individuals with an average age of 63.6 ± 7.5 years |
| Registration | Images are registered to a common MNI space for consistency |
| Resolution | 160x224x160 voxels with 1mm³ voxel size |
Summary
This presentation explores the use of Latent Diffusion Models (LDMs) for generating synthetic brain MRI scans, addressing key challenges in medical imaging:
- Purpose: Generate high-resolution MRI brain scans while handling data privacy and scarcity issues
- Method: Uses Latent Diffusion Models conditioned on age, sex, and brain structure volumes
- Dataset: UK Biobank T1-weighted MRI scans from healthy individuals (avg age 63.6 years)
- Results:
- Image quality 15x worse than real scans but 21x better than previous VAE-GAN approaches
- Diversity metrics comparable to real-world images
- Evaluated using Med3D-based FID and SSIM variants
- Technical Details:
- 160x224x160 voxel resolution
- 1mm³ voxel size
- MNI space registration
- Key Innovation: Uses DDIM sampling for faster training while maintaining quality
The work demonstrates significant progress in synthetic medical image generation, though limitations exist around dataset demographics and computational requirements.
Challenges & Limitations
- Training dataset contains only healthy individuals, limiting generalization to diseased populations. Moreover the average age of the training set is 63.6, further limiting the model’s ability.
- High memory requirements for training diffusion models.
- Semantic understanding of covariates remains complex.
References
This project was presented at MICCAI 2022 by Pinaya, Tudosiu, Dafflon et al.